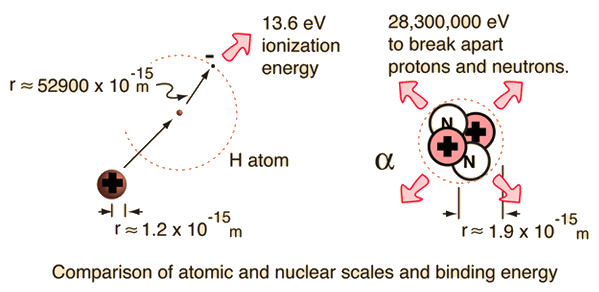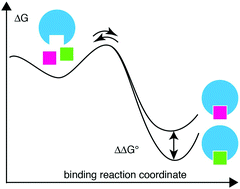
Mapping the energy landscape of protein–ligand binding via linear free energy relationships determined by protein NMR relaxation dispersion - RSC Chemical Biology (RSC Publishing)

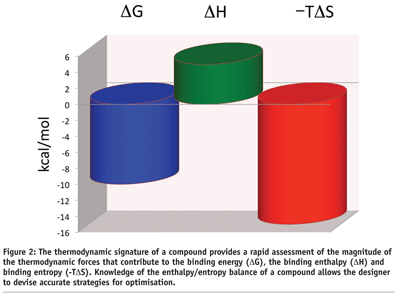
SOLVED: When a certain protein molecule binds to DNA, the entropy decreases by 2 kcal/degree: At the same time; it releases 700 kcal of enthalpy: What is the Gibbs energy change for

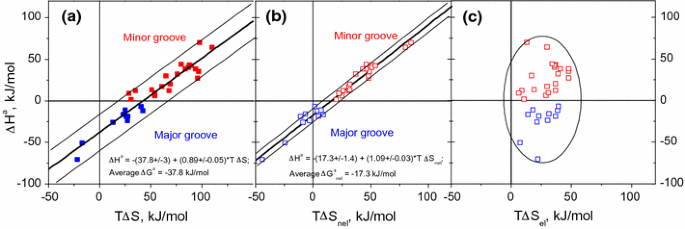
Left: Mean change in total enthalpy upon binding, hDH bind i, of each... | Download Scientific Diagram
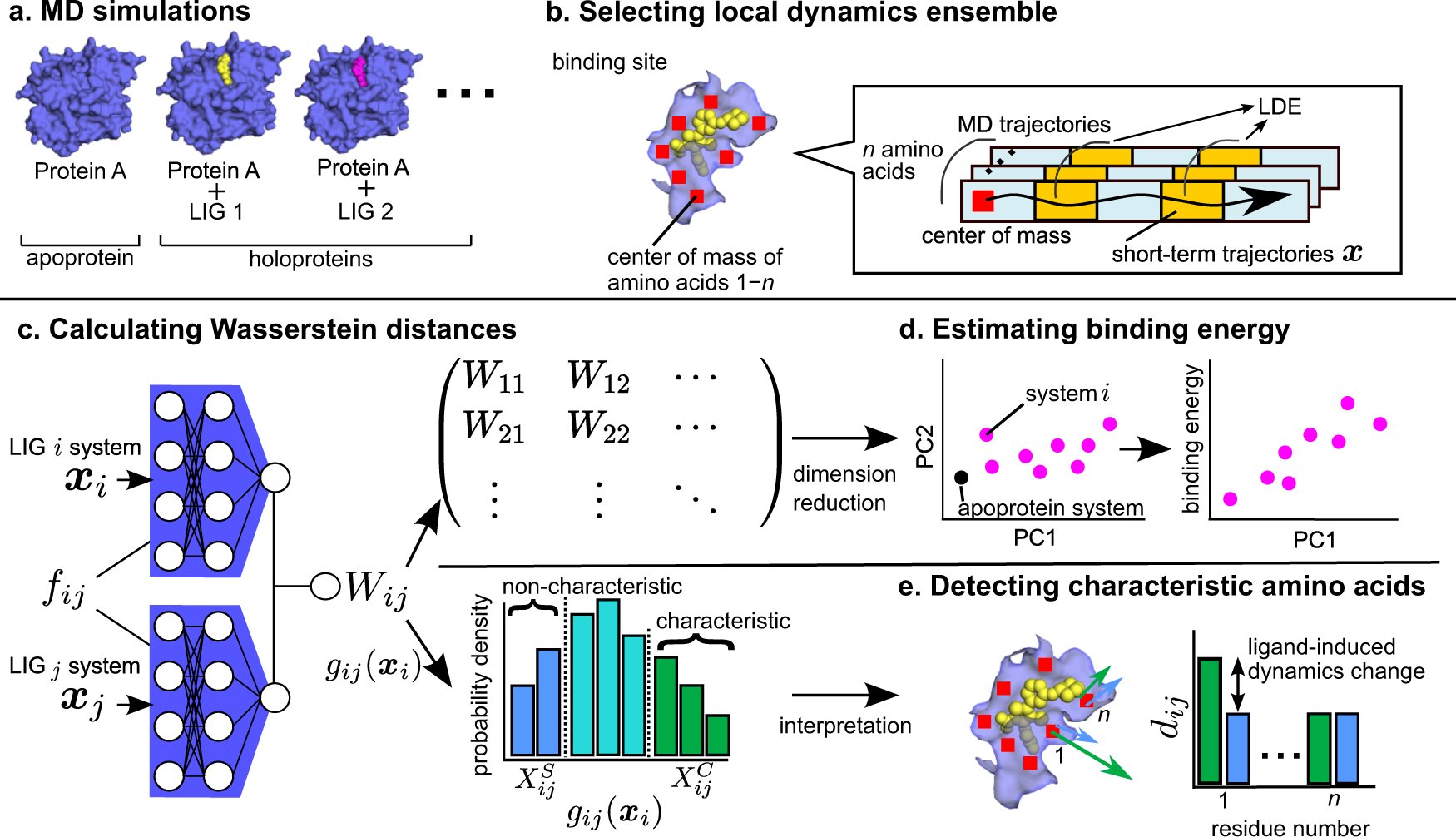
Differences in ligand-induced protein dynamics extracted from an unsupervised deep learning approach correlate with protein–ligand binding affinities | Communications Biology
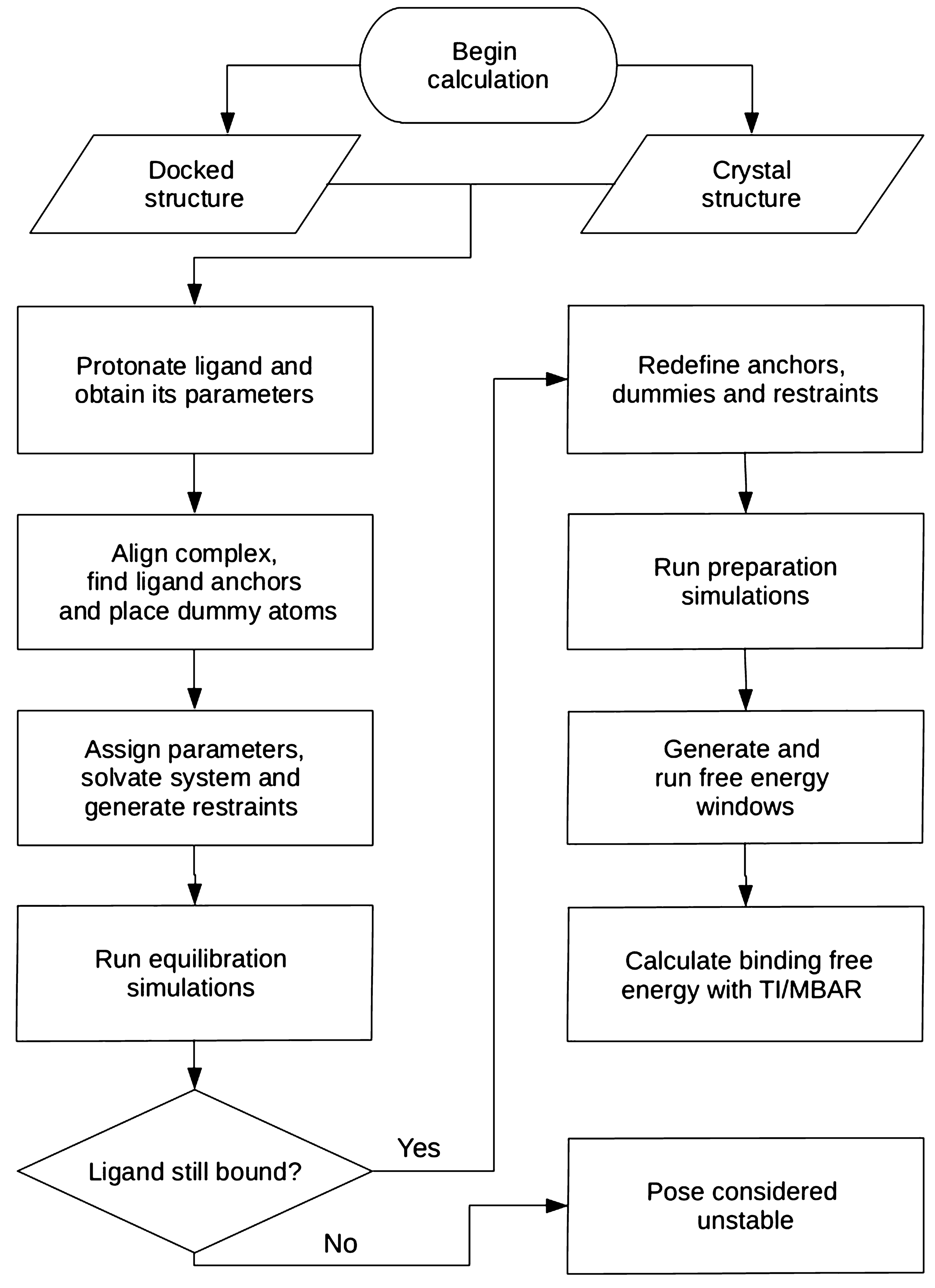
Automation of absolute protein-ligand binding free energy calculations for docking refinement and compound evaluation | Scientific Reports
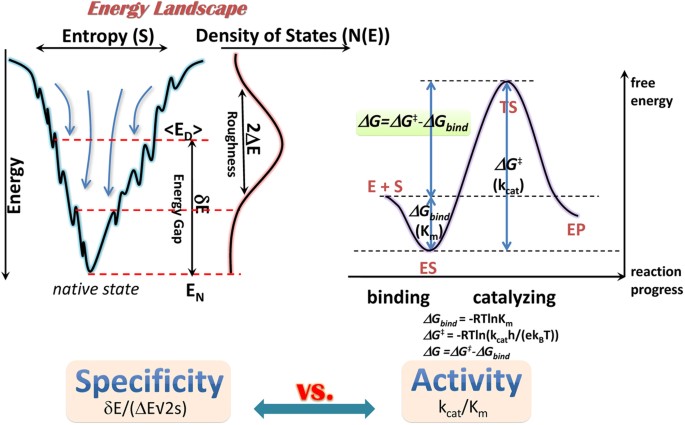
Energy Landscape Topography Reveals the Underlying Link Between Binding Specificity and Activity of Enzymes | Scientific Reports

Local potential energy: A novel QTAIM tool to quantify the binding energy of classical hydrogen bonds - ScienceDirect

Minimizing the Entropy Penalty for Ligand Binding: Lessons from the Molecular Recognition of the Histo Blood‐Group Antigens by Human Galectin‐3 - Gimeno - 2019 - Angewandte Chemie International Edition - Wiley Online Library