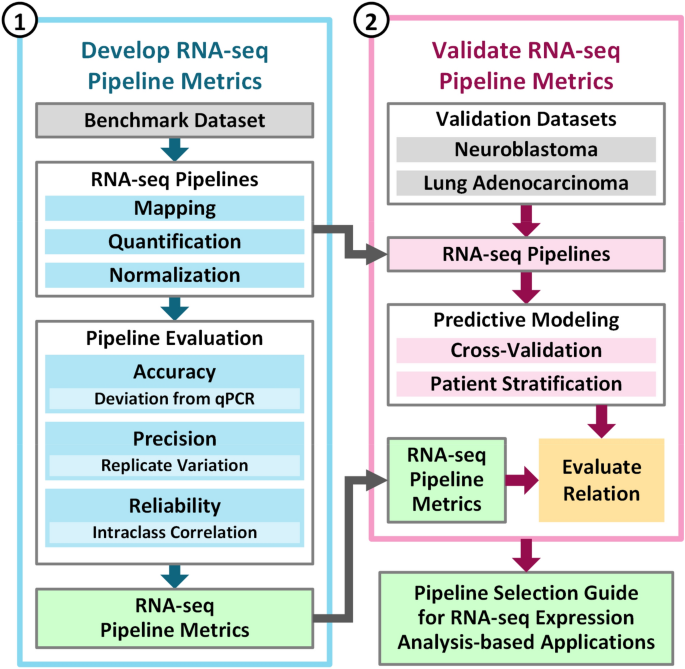
Impact of RNA-seq data analysis algorithms on gene expression estimation and downstream prediction | Scientific Reports

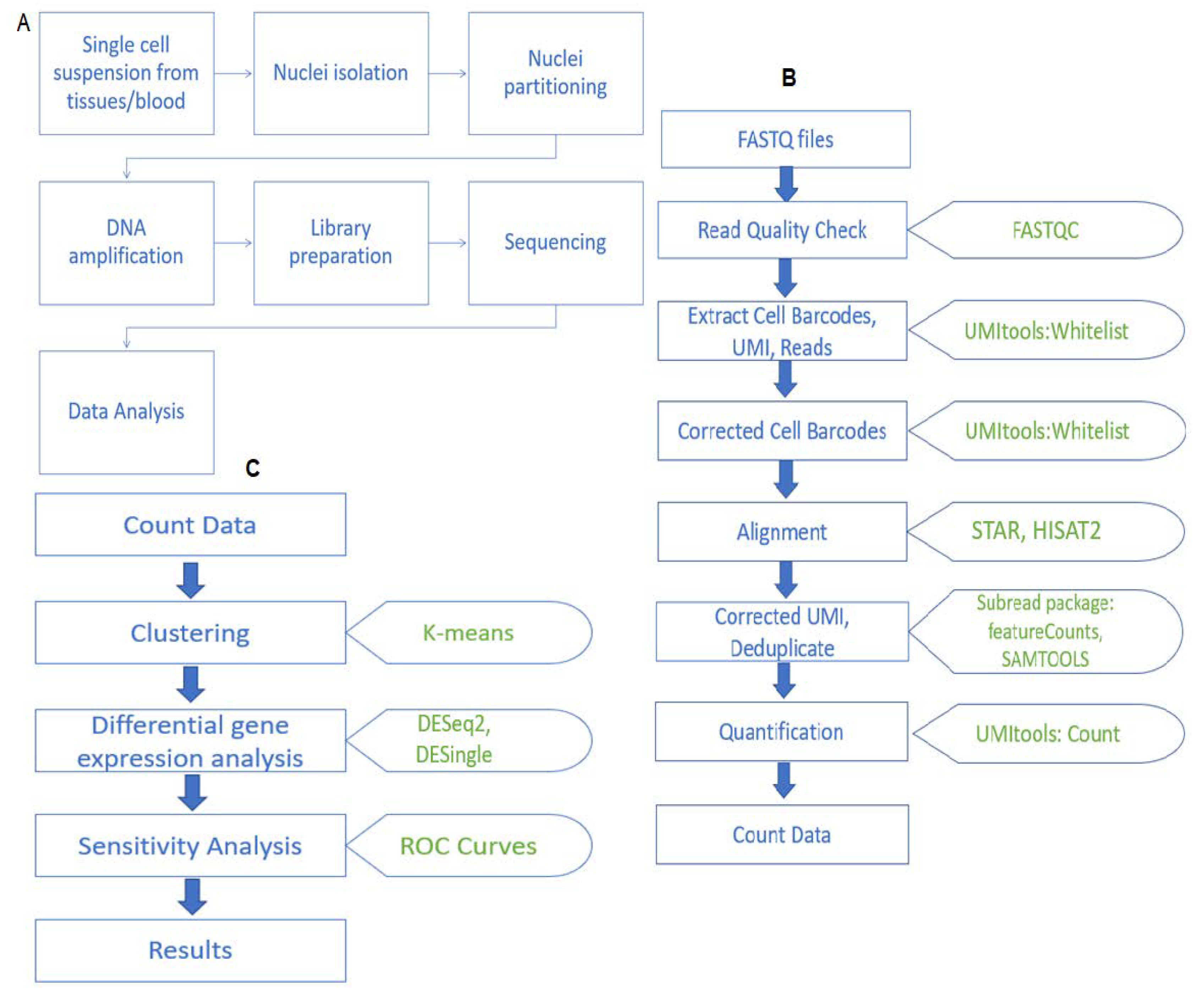
FNL collaborates with FDA to establish best practices for single-cell RNA sequencing | Frederick National Laboratory
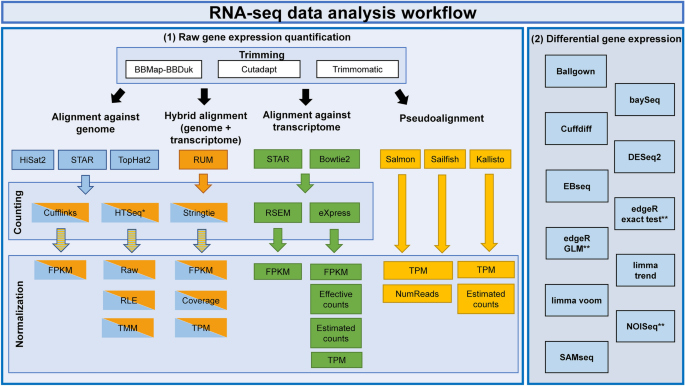
Systematic comparison and assessment of RNA-seq procedures for gene expression quantitative analysis | Scientific Reports

Amazon.com: RNA-seq Data Analysis: A Practical Approach (Chapman & Hall/CRC Computational Biology Series): 9781466595002: Korpelainen, Eija, Tuimala, Jarno, Somervuo, Panu, Huss, Mikael, Wong, Garry: Books

Guidelines for Setting Up a mRNA Sequencing Experiment and Best Practices for Bioinformatic Data Analysis | SpringerLink
Benchmarking RNA-seq differential expression analysis methods using spike-in and simulation data | PLOS ONE
Benchmarking RNA-seq differential expression analysis methods using spike-in and simulation data | PLOS ONE

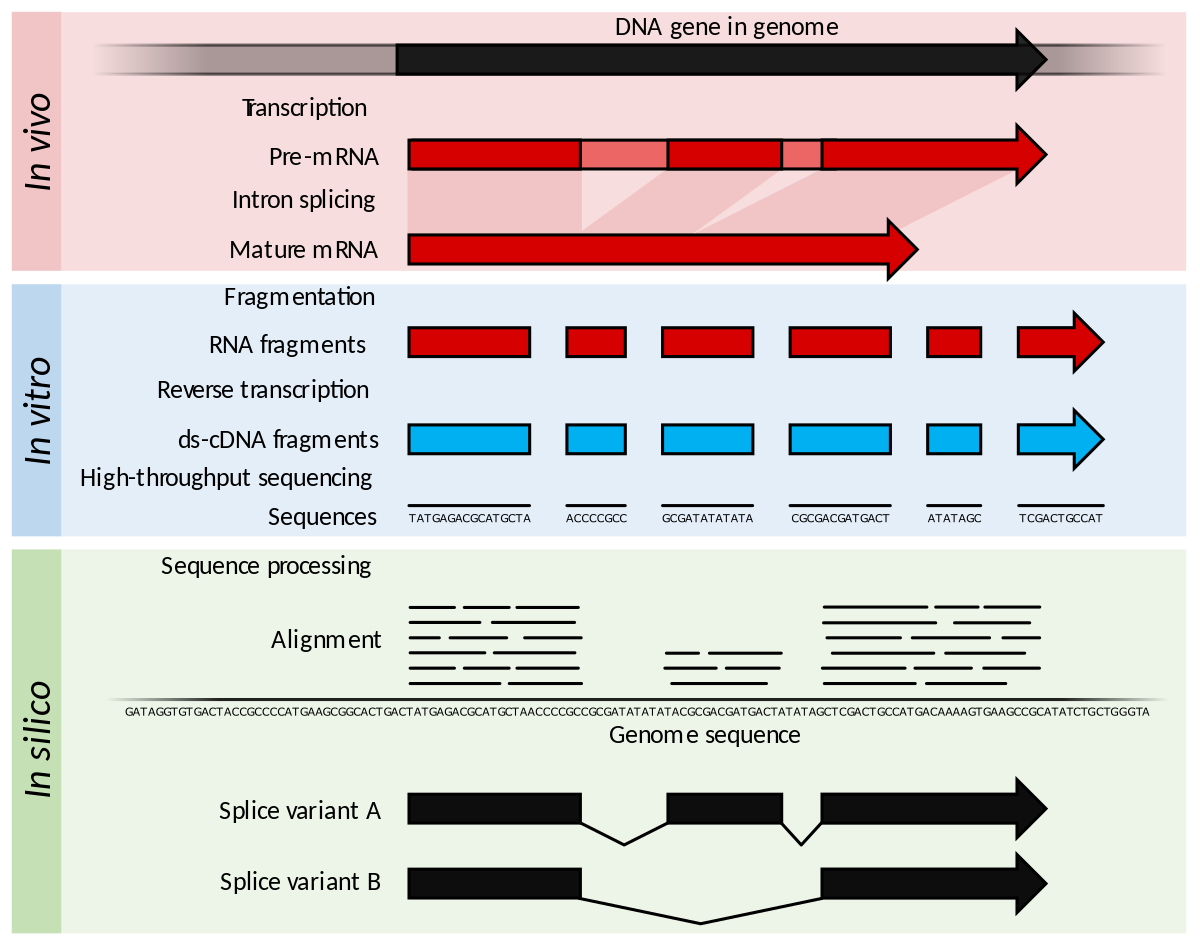



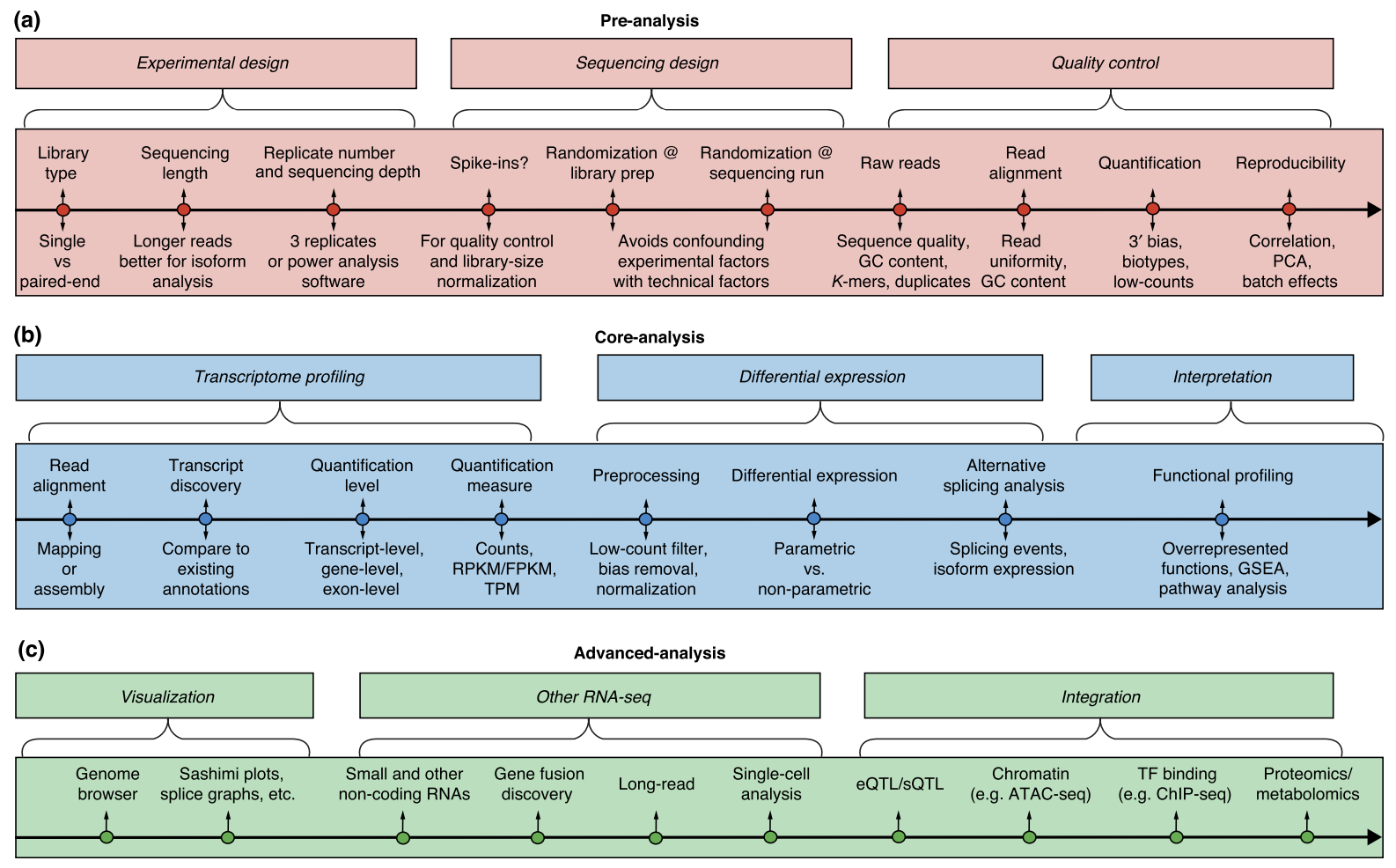
![PDF] RNA‐seq: Applications and Best Practices | Semantic Scholar PDF] RNA‐seq: Applications and Best Practices | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/54136f301fe8ae8ec0d5a67098c8587d728dfb23/4-Figure1-1.png)